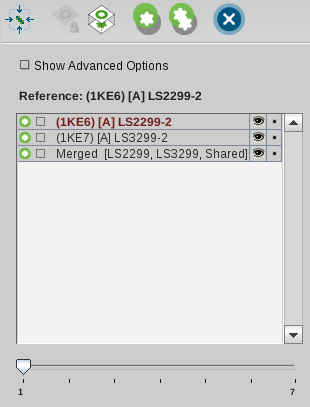
In case you would like to have more control over the
generation of shared feature pharmacophores you can use
LigandScout´s ability to edit existing pharmacophores in the
Alignment Panel
.
This includes aligning pharmacophores and
molecules by chemical features, merging these pharmacophores, and
interpolating their features to reduce overlaps. These editing
functionalities can be either performed fully automatically or
manually.
Automatically Merge Pharmacophores
The fully automatic way to merge pharmacophores involves first
aligning all of the selected elements, merging all features into a single
pharmacophore and interpolating features which overlap too much. To
perform this action select two or more elements from the Alignment List
and click on the icon
Generate Shared Feature Pharmacophore
.
LigandScout will then calculate several alignments. If there was
no valid alignment an error message will be displayed.
Otherwise, LigandScout will choose the best alignment and
use it for merging the pharmacophores of the selected elements.
Finally, features which overlap too much will be combined into a
single feature. After the algorithm has finished successfully, the
aligned, merged, and cleaned-up pharmacophore will be presented in
the Alignment List and the 2D/3D Viewer.
Manually Merge Pharmacophores
You can use the automatic alignment as described in
Aligning Pharmacophores and
Molecules
or you can align several structures manually by
selecting an element in the 3D Viewer, pressing Alt+Shift and using
LEFT
or
RIGHT
mouse button to change positions of this selected
element relative to the other elements.
Two or more pharmacophores can be merged by selecting two or
more elements in the Alignment List and pressing CTRL+ALT+M
(Option+Command+M on MAC). The resulting pharmacophore will contain
a copy of every feature from the selected pharmacophores or the
pharmacophores of the selected molecules. The resulting
pharmacophore can be cleaned up from overlaps automatically
(see below).
Interpolate All Features Within Tolerance
Select one pharmacophore in the Alignment List and press
CTRL+ALT+T (Option+Command+T on MAC). This will trigger LigandScout
to detect and group overlapping features automatically. Each group
of features will be replaced by the interpolated feature. So the
resulting pharmacophore will contain minimal overlap.
You can also select features of the same type and from the
same pharmacophore in the 3D Viewer and have them interpolated by
pressing CTRL+E (Command+E on MAC). Here, the restrictions are not
as tight as in the automatic way. You can select features within
the range of five times the average tolerance of the selected
features. If all possible pairs formed from the selected features
are within this range this interpolation will be performed. The
previously selected features will be replaced by the interpolated
ones.
Combine Features in One Pharmacophore
LigandScout supports the combination of all features of
several aligned pharmacophores to one model. This is useful, e.g., if
there are two or protein-ligand complexes available and you want to
combine the interactions formed by these ligands. Use the
>
command to perform the
automated combination of the currently selected
pharmacophores.
In most cases you may want to downsize the number of features
of the new model, especially, if there are features of the same
type adjacent to each other. Therefore, you can either use the
>
command
for a global merge, or you manually select the features (of the
same feature type) that you want to interpolate and use the
>
command for manual merging (see above).