|
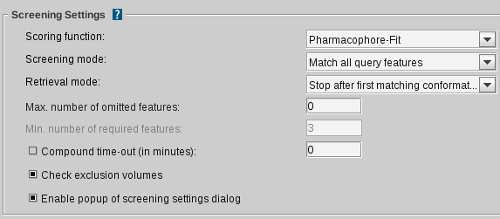
Scoring function
|
Choose the scoring function type that validates the
alignments. The Pharmacophore-Fit scoring function only considers
pharmacophoric features and the feature RMSD. The Relative Pharmacophore-Fit
scores the number of matching pharmacophore features and the RMSD
of the pharmacophore alignment normalized to [0..1]. (default: Pharmacophore-Fit)
|
|
Screening mode
|
In Match all query features mode the database is
browsed to find molecules that are fully matching the given set of query
pharmacophores. Whereas in Fragment screening mode, the database
molecules are fitted into the query pharmacophores. In the latter case,
the query is partially matched. (default: Match all query features)
|
|
Retrieval mode
|
Stopping criteria per compound. Stop after first
matching conformation: the screening for a molecule stops when the first
matched conformation is found. Get best matching conformation: all
conformations are examined for the best matching conformation. Get all
matching conformation: all conformations that match the query pharmacophore
are shown as screening hits. The last two modes could slow down screening.
(default: Stop after first matching conformation)
|
|
Max. num. of omitted features
|
Defines the maximum number of query features that can be
omitted during the Match all query features mode.
If zero is specified, the whole query pharmacophore
must be matched to be a valid hit.
(default: 0)
|
|
Min. num. of required features
|
Specifies the minimum number of required features that has to match between the
library molecule and the query pharmacophore to be a valid hit.
This setting is only enabled if the Fragment screening mode is chosen.
(default: 3)
|
|
Compound time-out (in minutes)
|
Enables the maximum screening time to spend per compound. (default: off)
|
|
Check exclusion volumes
|
Enables the consideration of exclusion volumes
during the screening process. (default: enabled)
|
|
Enable popup of screening settings dialog
|
Enables the settings dialog of the screening process. (default: enabled)
|