LigandScout supports commonly used file formats as well as
its own file formats to exchange information. As the main focus of LigandScout
is drug development and pharmacophore modeling, all file formats either
deal with molecular data, pharmacophore models, or display-related
information.
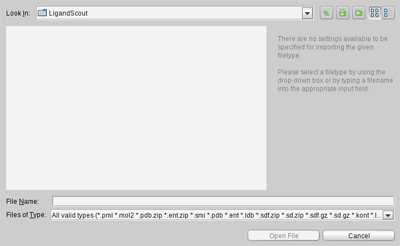
The
File
menu provides two
possibilities to import data: the
Open
command (CTRL+O) and the
Insert
command (CTRL+I). Both trigger
the display of the file dialog that facilitates the import of files
that are being loaded into the current view,
whereas the former clears the view first.
The supported import file formats can be differentiated into
LigandScout, molecule and other file formats.
The following table
gives an overview of the file formats including
name and file extension list.
| File Format | Name | Extension(s) |
|---|---|---|
| LigandScout | Library in Local Database | .ldb |
| Ligand Set Description | .lsd | |
| Pharmaceutical Markup Language | .pml | |
| Compressed Pharmaceutical Markup Language | .pmz | |
| Session | .ses | |
| Molecule | MDL Molfile | .mol |
| Tripos Sybyl MOL2 Molecule | .mol2 | |
| MDL Structure-Data | .sd .sdf | |
| GZip Compressed MDL Structure-Data | .sd.gz .sdf.gz | |
| Zip Compressed MDL Structure-Data | .sd.zip .sdf.zip | |
| Daylight Chemical SMILES String | .smi | |
| Other | Protein Data Bank (PDB) | .pdb .ent |
| GZip Compressed Protein Data Bank (PDB) | .pdb.gz .ent.gz | |
| Zip Compressed Protein Data Bank (PDB) | .pdb.zip .ent.zip | |
| Molecular Discovery GRID | .kont |
Table 3.1. Supported import file formats
The PDB file format provides 3D information
about the macromolecule from X-ray or NMR sources. This data plays an important
role in deriving a pharmacophore model representing the bound state of a ligand inside a
protein binding pocket. The Molfile characterizes a single molecular structure
whereas SD(F) and MOL2 files can store more than one molecule (molecule library).
In LigandScout, a single molecule can be used to create a pharmacophore model or
to align to other molecules/pharmacophores. The latter allows LigandScout
to import compound collections. For instance investigations of these
ligands with respect to their protein environment, and also for generating
pharmacophores based on docking poses. Moreover, LigandScout can
filter molecules from a molecule library that fulfill a desired pharmacophoric
pattern. Compared to the other file formats, SMILES files represent only the 2D
depiction of the molecules.
The
2D View
of LigandScout uses the
2D information of the ligand to visualize the derived pharmacophoric features.
The PML file format can be stored and opened with different combinations of content elements,
for instance a pharmacophore of a ligand-protein complex or alignments of molecules.
The PMZ format contains the compressed PML data. The LSD file format
is used in the
Ligand-Based Modeling Perspective
to generate pharmacophores from a set of ligands. The LDB file format are used to
store/load molecule libraries. This can be useful for investigations of
compounds with respect to their protein environment, and also for generating
pharmacophores based on docking poses. For more information on advanced export settings,
see also
the section called “Advanced Export Settings”
.
The GRID Map file format can be imported to LigandScout
to analyze energetically favorable regions in the binding pocket of a protein.
For more information about this topic, please see
the section called “GRID Maps Visualization”
.
LigandScout supports data export in several chemical and
graphical file formats. Use the command (CTRL+SHIFT+S) in the menu
File
>
Save as File
which enables you to
save your macromolecule, ligand modifications, and pharmacophore models.
A session can be stored in the menu
File
>
Sessions
>
Save as File
.
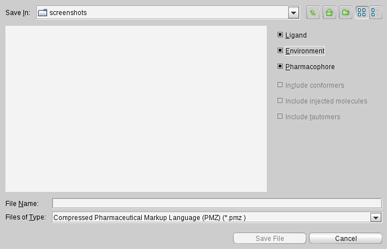
Depending on the selected file format and content, the
Save File
dialog provides
advanced export settings
.
The following table
gives you a summary of the supported export file formats together with which data
of your work that can be exported in a particular format.
Each format facilitates the export of certain information:
molecules (M), ligand environment (E), pharmacophore models (P), and
the current content of either 3D View or 2D View (V).
| File Format | Name | Extension(s) | M | E | P | V |
|---|---|---|---|---|---|---|
| LigandScout | Compressed Pharmaceutical Markup Language | .pmz | x | x | x | |
| Pharmaceutical Markup Language | .pml | x | x | x | ||
| Ligand Set Description | .lsd | x | x | |||
| Library in Local Database | .ldb | x | ||||
| Session | .ses | x | x | x | ||
| Molecule | Daylight Chemical Smiles String | .smi | x | |||
| MDL Structure-Data | .sd .sdf | x | ||||
| GZip Compressed MDL Structure-Data | .sd.gz .sdf.gz | x | ||||
| Zip Compressed MDL Structure-Data | .sd.zip .sdf.zip | x | ||||
| Maestro Molecule | .mae | x | ||||
| MDL Molfile | .mol | x | ||||
| Tripos Sybyl MOL2 Molecule | .mol2 | x | ||||
| Pharmacophore | Catalyst Hypoedit Script | .hypoedit | x | |||
| Fred 2 Constraint Map | .fred2map | x | ||||
| Phase Files | .xyz .xvol .def .mae | x | ||||
| CCG MOE Format | .ph4 | x | ||||
| Image | Encapsulated PostScript | .eps | x | |||
| Graphics Interchange Format | .gif | x | ||||
| Joint Photographers Expert Group Format | .jpg | x | ||||
| MacroMedia Flash File Format | .swf | x | ||||
| POV-Ray Scene Desription | .pov | x | ||||
| Portable Document Format | x | |||||
| Portable Network Graphics Format | .png | x | ||||
| PostScript | .ps | x | ||||
| RAW Image | .raw | x | ||||
| Scalable Vector Graphics | .svg | x | ||||
| GZip Compressed Scalable Vector Graphics | .svgz | x | ||||
| UNIX Portable PixMap | .ppm | x | ||||
| Windows Enhanced Metafile | .emf | x | ||||
| Other | Protein Data Bank (PDB) | .pdb .ent | x | x | ||
| GZip Compressed Protein Data Bank (PDB) | .pdb.gz .ent.gz | x | x | |||
| Zip Compressed Protein Data Bank (PDB) | .pdb.zip .ent.zip | x | x |
Table 3.2. Supported export file formats
LigandScout has its own file format, called PML. The
advantage of this format is that the user can select the available
export content individually or in combination (ligand, environment,
pharmacophore, conformers, injected molecules, and tautomers).
LigandScout offers you the possibility to export LigandScout's pharmacophore
models so that you can use them in other external programs
like Catalyst, MOE and Phase (see
the section called “Preparing a Pharmacophore for Virtual Screening with External Applications”
).
You can save your content as PDB or ENT file format if you want to
export the ligand, environment and include hydrogen information.
The SMILES string of molecules can also be exported as SMI files. You can include
in the files the canonical order of ligand atoms and/or stereo chemical properties.
Single molecules can be stored in mol files, as 2D or 3D data. Additionally,
stereo-chemical properties can be saved optionally.
The storage of a molecule
library in LDB, MOL(2) and SD(F)
files can be customized. You can export a
Complete library
or a part of it.
A subset of a library can be chosen by
filtering your library
according to a property (e.g. by an occurring name), marking molecules or
by visible molecules only.
If you want to export the properties of your library table, choose the
Write properties
option.
The MOL2 and SD(F) file formats
offer you additionally the possibility to
Write all conformations
or
Write 2D/3D coordinates
of the (selected)
molecules in the library.
If you want to export currently
displayed content as an image file (e.g. gif, svg, etc.)
you can choose between the 2D or 3D View. The 2D content can be saved as it
is by the
Keep current layout
or you can automatically
store it as
horizontal/vertical layout
. After
a screening run, you can export the
ROC plot
as image.
The image size can be set in both 2D and 3D.